热点招聘
高校推荐
南开大学面向全球诚聘英才
中国政法大学2019年度教学科研岗位补充招聘启事
中国劳动关系学院2019年招聘简章
中国青年政治学院2019年教师岗位招聘公告
美国亚利桑那大学药理系招聘做膜片钳博后
浙江大学诚邀海外优秀青年人才
武汉大学2019年招聘
首都医科大学2019年公开招聘公告
首都师范大学诚聘高层次国际人才
北京外国语大学高层次人才引进公告
中国人民大学2019学年教师岗位招聘公告
新加坡国立大学 Prof Ouyang Jianyong 博后招聘
同济大学第三届“国际青年学者论坛”
香港理工大学 招聘无人机视觉控制方向 全奖博士后
中国财政发展协同创新中心2019年公开招聘海外留学人...
美国梅奥医学中心核酸分子生物学方向博后招聘
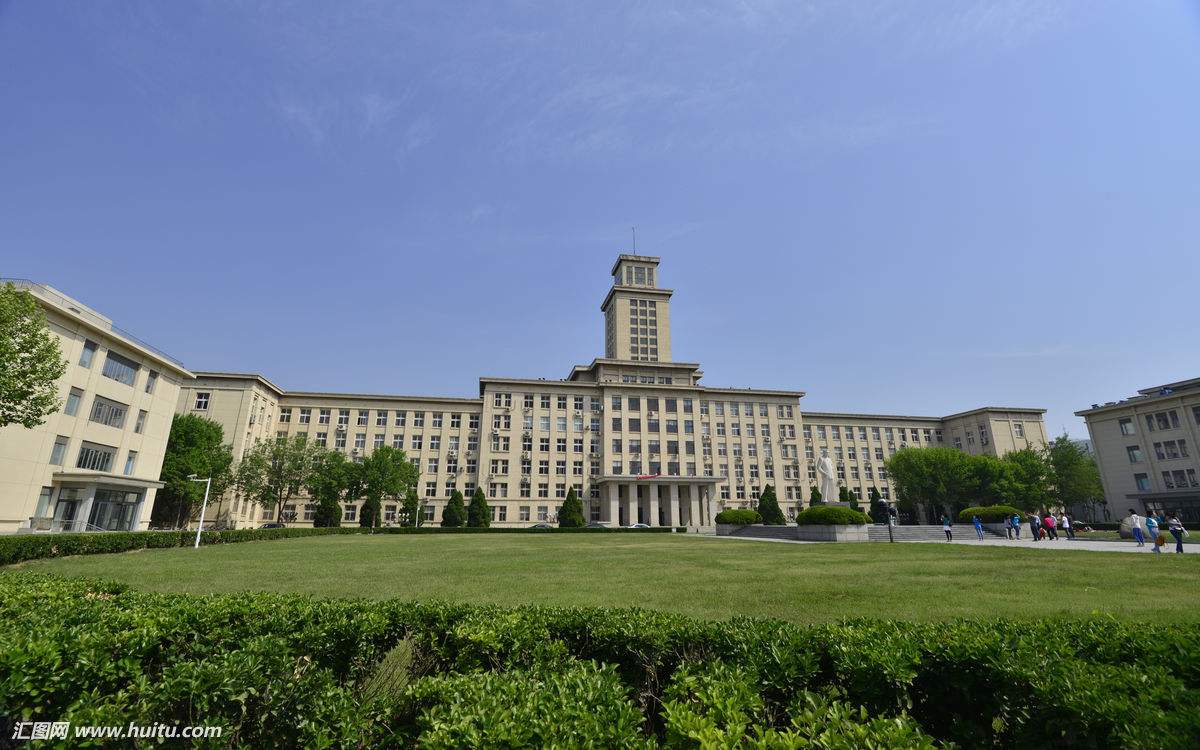 南开大学面向全球诚聘英才
南开大学面向全球诚聘英才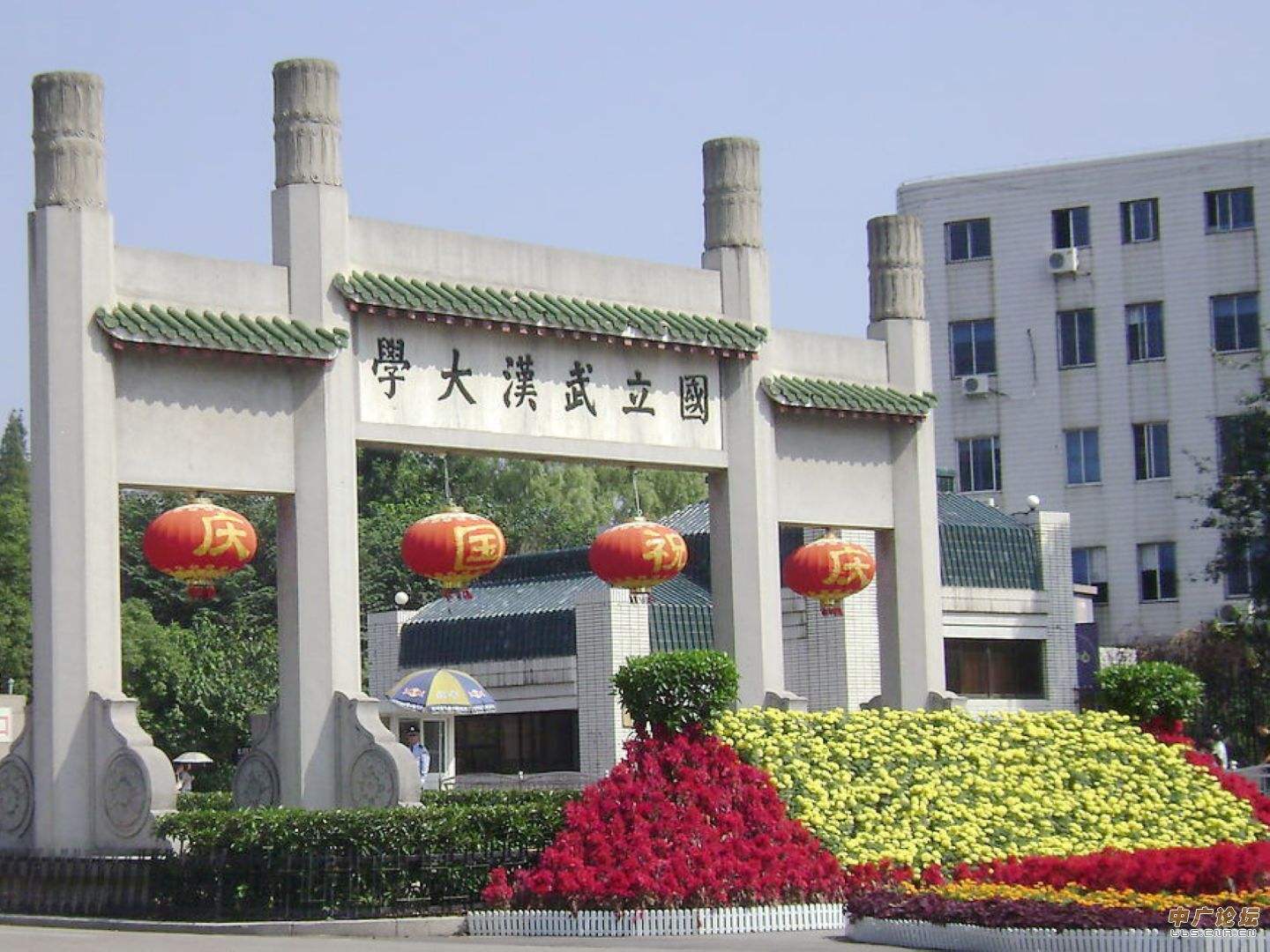 武汉大学2019年招聘
武汉大学2019年招聘 国际关系学院2019年管理教辅人员招聘启事
国际关系学院2019年管理教辅人员招聘启事 南洋理工大学 招收 计算机视觉(图像视频挖掘) 博士后2名
南洋理工大学 招收 计算机视觉(图像视频挖掘) 博士后2名 澳门大学电化学博后招聘
澳门大学电化学博后招聘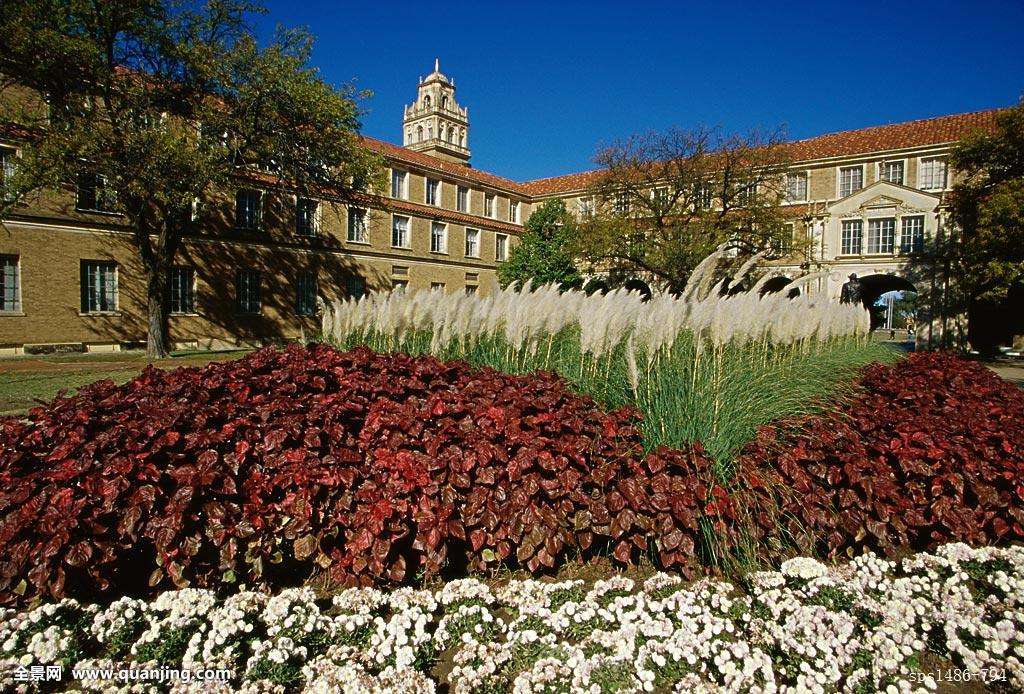 美国德克萨斯大学招聘材料生物博后
美国德克萨斯大学招聘材料生物博后 中国农业大学2019年人才招聘启事
中国农业大学2019年人才招聘启事 中国人民大学2019学年教师岗位招聘公告
中国人民大学2019学年教师岗位招聘公告 美国密苏里科技大学博士后招聘
美国密苏里科技大学博士后招聘 宾夕法尼亚州立大学招收博后
宾夕法尼亚州立大学招收博后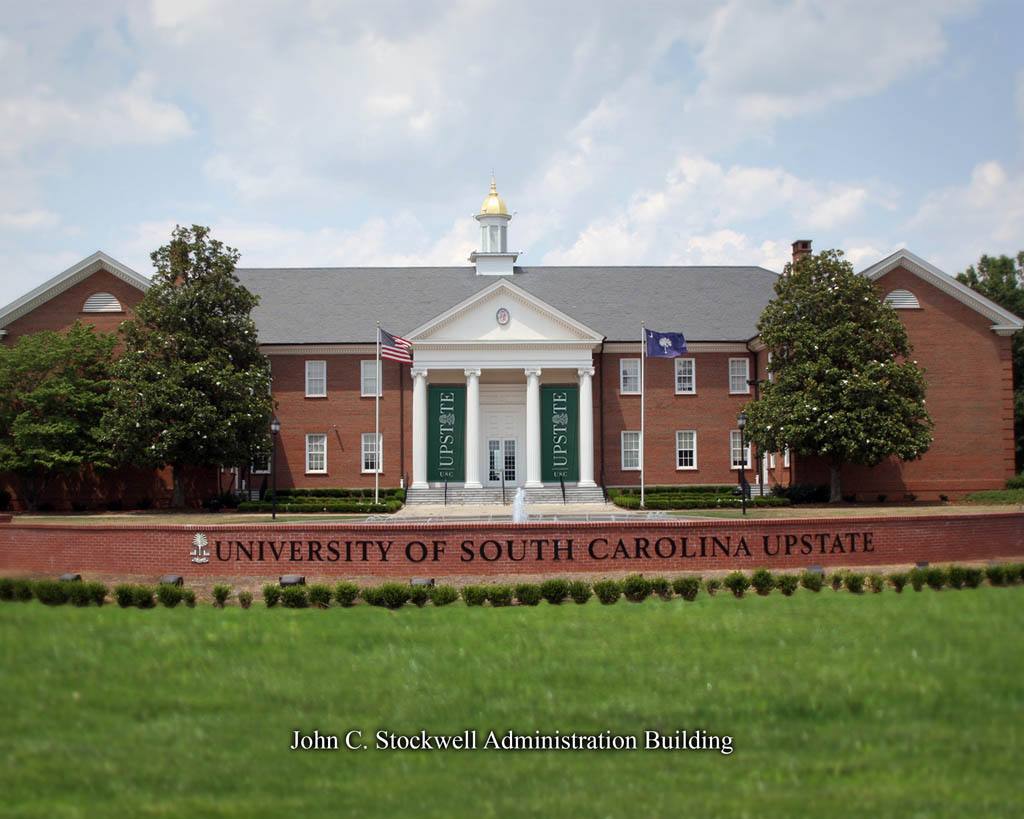 美国南卡罗来纳大学招聘药物输送, 生物材料,神经科学方向博士后
美国南卡罗来纳大学招聘药物输送, 生物材料,神经科学方向博士后 外交学院博士后招聘启事
外交学院博士后招聘启事
高校引才
中央民族大学2019年教学科研岗位应届高校毕业生公开...
香港理工大学招聘无人机视觉控制方向博士后
南开大学面向全球诚聘英才
中国人民大学2019学年教师岗位招聘公告
康奈尔大学王飞教授招博后
中央财经大学绿色金融国际研究院招聘启事
同济大学第三届“国际青年学者论坛”
武汉大学2019年招聘
美国德克萨斯大学招聘材料生物博后
南京大学2019年高层次人才招聘
北京语言大学2019年海内外优秀人才招聘启事
美国内布拉斯加大学林肯分校招收有机合成方向博后
哈佛大学汪淏田课题组诚招博士后
首都医科大学2019年公开招聘公告
中国政法大学2019年度教学科研岗位补充招聘启事
北京舞蹈学院2019年招聘公告
 佐治亚大学诚聘具有化学、材料、物理和生物等相关背景的博士后
佐治亚大学诚聘具有化学、材料、物理和生物等相关背景的博士后 中国劳动关系学院2019年招聘简章
中国劳动关系学院2019年招聘简章 北京外国语大学高层次人才引进公告
北京外国语大学高层次人才引进公告 南开大学面向全球诚聘英才
南开大学面向全球诚聘英才 首都医科大学2019年公开招聘公告(第五批)
首都医科大学2019年公开招聘公告(第五批) 中国科学院大学教学科研岗位人才招聘
中国科学院大学教学科研岗位人才招聘 中国农业大学2019年人才招聘启事
中国农业大学2019年人才招聘启事 中国青年政治学院2019年教师岗位招聘公告
中国青年政治学院2019年教师岗位招聘公告 中国人民大学2019学年教师岗位招聘公告
中国人民大学2019学年教师岗位招聘公告 北京舞蹈学院2019年招聘公告
北京舞蹈学院2019年招聘公告 首都师范大学诚聘高层次国际人才
首都师范大学诚聘高层次国际人才 同济大学第三届“国际青年学者论坛”
同济大学第三届“国际青年学者论坛” 武汉大学2020招聘人才
武汉大学2020招聘人才 中国政法大学2019年度教学科研岗位补充招聘启事
中国政法大学2019年度教学科研岗位补充招聘启事 中国财政发展协同创新中心2019年公开招聘海外留学人才公告
中国财政发展协同创新中心2019年公开招聘海外留学人才公告 加州大学河滨分校博士后招聘
加州大学河滨分校博士后招聘 东南大学2019年人才需求
东南大学2019年人才需求 美国亚利桑那大学药理系招聘做膜片钳博后
美国亚利桑那大学药理系招聘做膜片钳博后 耶鲁大学实验室招收博士后
耶鲁大学实验室招收博士后 波士顿大学-Doctor. Yi 课题组招收博后
波士顿大学-Doctor. Yi 课题组招收博后 中央民族大学2019年教学科研岗位应届高校毕业生公开招聘公告
中央民族大学2019年教学科研岗位应届高校毕业生公开招聘公告 首都医科大学2019年公开招聘公告(第五批)
首都医科大学2019年公开招聘公告(第五批) 澳门大学电化学博后招聘
澳门大学电化学博后招聘 宾夕法尼亚州立大学招收博后
宾夕法尼亚州立大学招收博后 澳大利亚麦考瑞大学博士后招聘2019年面向海内外招聘优秀生物医学专业的博士后
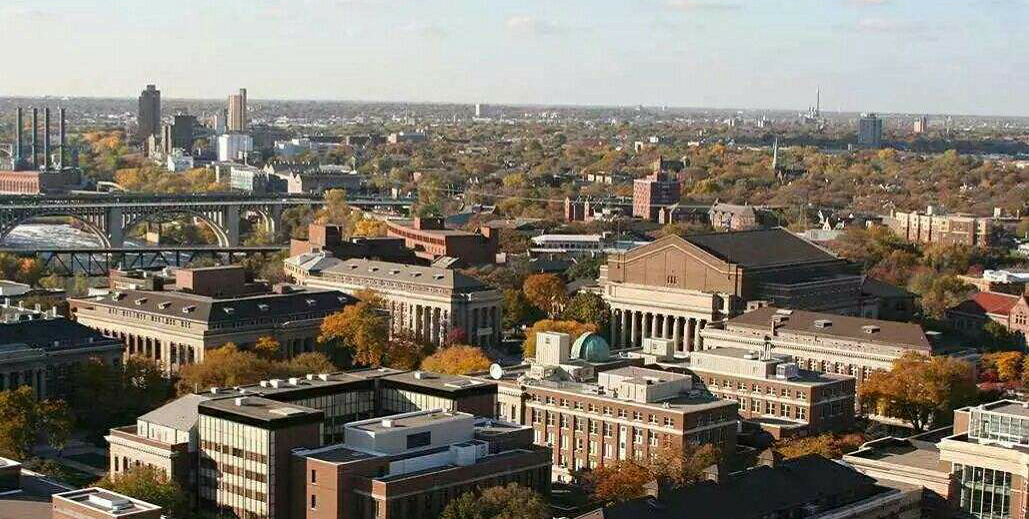
澳大利亚麦考瑞大学博士后招聘2019年面向海内外招聘优秀生物医学专业的博士后 美国明尼苏达大学招聘生物信息学博后
美国明尼苏达大学招聘生物信息学博后 美国纽约西奈山医学院招聘生命衰老学计算-生物信息方向博士后
美国纽约西奈山医学院招聘生命衰老学计算-生物信息方向博士后 南洋理工大学招收计算机视觉(图像视频挖掘)博士后2名
南洋理工大学招收计算机视觉(图像视频挖掘)博士后2名 新加坡国立大学 Prof Ouyang Jianyong 博后招聘
新加坡国立大学 Prof Ouyang Jianyong 博后招聘 南京大学2019年高层次人才招聘
南京大学2019年高层次人才招聘 上海交通大学感知与导航研究所博士后招聘
上海交通大学感知与导航研究所博士后招聘 北京语言大学2019年海内外优秀人才招聘启事
北京语言大学2019年海内外优秀人才招聘启事 美国马里兰大学医学院招博士后 (有IPSC 和 Sequencing 经验的优先考虑)
美国马里兰大学医学院招博士后 (有IPSC 和 Sequencing 经验的优先考虑) 美国南佛罗里达大学李霄鹏课题组拟招化学材料博士后
美国南佛罗里达大学李霄鹏课题组拟招化学材料博士后 圣路易斯华盛顿大学招聘固态锂金属电池方向博士后
圣路易斯华盛顿大学招聘固态锂金属电池方向博士后 美国密苏里科技大学博士后招聘
美国密苏里科技大学博士后招聘 北京电影学院2019年度博士后招收简章
北京电影学院2019年度博士后招收简章 美国内布拉斯加大学林肯分校招收有机合成方向博后
美国内布拉斯加大学林肯分校招收有机合成方向博后 香港理工大学招聘无人机视觉控制方向 全奖博士后研究员或者研究助理,访问学者等
香港理工大学招聘无人机视觉控制方向 全奖博士后研究员或者研究助理,访问学者等 国际关系学院2019年管理教辅人员招聘启事
国际关系学院2019年管理教辅人员招聘启事 美国南卡罗来纳大学招聘药物输送, 生物材料,神经科学方向博士后
美国南卡罗来纳大学招聘药物输送, 生物材料,神经科学方向博士后 北京化工大学2019年管理人员招聘启事
北京化工大学2019年管理人员招聘启事 美国德克萨斯大学招聘材料生物博后
美国德克萨斯大学招聘材料生物博后 外交学院博士后招聘启事
外交学院博士后招聘启事 浙江大学面向校外公开招聘“求是特聘学者”启事
浙江大学面向校外公开招聘“求是特聘学者”启事 美国犹他州立大学招聘博士后 微流控 微工程 Utah State Univ
美国犹他州立大学招聘博士后 微流控 微工程 Utah State Univ 哈佛大学汪淏田课题组诚招博士后
哈佛大学汪淏田课题组诚招博士后 美国休斯顿大学博后招聘
美国休斯顿大学博后招聘
高校展示
美国纽约西奈山医学院招聘生命衰老学计算-生物信息方...
北京化工大学2019年管理人员招聘启事
外交学院博士后招聘启事
美国爱荷华州立大学Xu Jing教授课题组博后和博士招聘...
澳门大学电化学博后招聘
首都师范大学诚聘高层次国际人才
中国农业大学2019年人才招聘启事
浙江大学面向校外公开招聘“求是特聘学者”启事
北京外国语大学高层次人才引进公告